Abstract
The study was carried out with an aim to isolate, characterize and determine the antibiotics resistance pattern of bacteria isolated from a tertiary hospital in Abakaliki. Waste water effluents were collected from various units of the Alex Ekwueme University Teaching Hospital, Abakaliki and analysed at the Applied Microbiology Laboratory unit of the Ebonyi State University Abakaliki using standard microbiology techniques. Susceptibility of the bacterial isolates to various classes of antibiotics was determined by disc diffusion method. Production of Extended spectrum beta lactamase was determined by the double disc synergy method and the presence methicillin resistant Staphylococcus aureus was determined using Cefoxitin and methicillin disc diffusion method. Our findings showed that Escherichia coli had the highest frequency of bacterial occurence (43.24%) followed by S. aureus (40.54%), Klebsiella (8.11%) and Campylobacter (8.11%). Waste water effluents from the ward complex had the highest distribution of bacteria isolates (40.54%) followed by effluents from the laboratory complex. Effluents from the theatre had the least distribution (2.70%). Out of 15 S. aureus isolates obtained from this study, 46.66% (7) were MRSA positive while out of 16 E. coli isolated from this study, 31.25% (5) were positive for ESBL production. Multi-antibiotic resistance index of S. aureus showed the highest values to be from the ward complex (1.00) and laboratory complex (0.29). Multi-antibiotic resistance index of Campylobacter species showed the highest values to be from the ward complex (0.75) and the least from laboratory complex (0.50) and that of Klebsiella species showed the highest values to be from accident/emergency (0.88) and the least from the ward complex (0.63). Multi-antibiotic resistance index of E. coli showed the highest values to be from the ward complex and laboratory complex (1.00). The presence of antimicrobial resistant bacteria from hospital wastewaters showed the spreading of AMR bacteria into the environmental through wastewater. The presence of high MDR bacteria in hospital wastewater may impose public health challenges because they can transmit resistance traits to other enteric pathogenic bacteria in the community.
|
Published in
|
Biomedical Sciences (Volume 12, Issue 2)
|
|
DOI
|
10.11648/j.bs.20261202.11
|
|
Page(s)
|
26-33 |
|
Creative Commons
|

This is an Open Access article, distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution and reproduction in any medium or format, provided the original work is properly cited.
|
|
Copyright
|
Copyright © The Author(s), 2026. Published by Science Publishing Group
|
Keywords
Hospital, Waste, Water, Antibiotics, Resistant, Abakaliki
1. Introduction
Hospital waste is categorized into pathological waste; body fluids from surgery, infectious waste from laboratories, pharmaceutical waste; out-of-date pharmaceutical products, and chemical wastes; used solvents, disinfectants, pesticides, and diagnostic chemicals, aerosol containers and gas, and open sources used in
in vitro diagnosis or nuclear medical therapy
| [4] | Dadi, B. R., Girma, E., Tesfaye, M and Seid, M (2021). Assessment of the Bacteriological Profile and Antibiotic Susceptibility Patterns of Wastewater in Health Facilities of Ethiopia. International Journal of Microbiology, 9(6): 45-50. |
[4]
. Sewage from hospitals, usually referred to as hospital waste water, is defined as a special category of waste which comprises all wastes, biological or non-biological, that are discarded from hospitals/healthcare centers and not intended for further use
| [11] | Odu, C. E., Egbere, J. O., Onyimba, I. A., Ghamba, P. E., Godiya, S., Isyaka, T. M., Collins- Odu, J. O., Idigo, M. A and Ngene, A. C. (2022). Resistance Profiles of Bacteria Isolated from Wastewater in the State Specialist Hospital Maiduguri. European Journal of Biology and Biotechnology, 3(3), 1–6. |
| [12] | Oyeleke, S., Istifanus, N and Manga, S (2008). The effects of hospital solid waste on the receiving environment. International Journal of Integrative Biology, 3(3): 191. |
[11, 12]
.
The important usage of water in hospitals gives significant volumes of waste loaded with microorganisms (the majority of which being pathogenic), heavy metals, toxic chemicals, disinfectant and radioactive elements
| [10] | Kummerer, K (2004). Resistance in the environment. Journal of Antimicrobial Chemotherapy, 54(2): 311–320. |
[10]
.
However, many non-metabolized drugs excreted from patients and residual disinfectants enter into wastewater, which finally interacts with the microflora of hospital sewage. Hospital wastewater are considered and best described as hot spots for the dissemination of antibiotic and disinfectant resistance bacteria that could threaten public health upon water reuse
| [9] | Iweriebor, B. C., Obi, L. C and Okoh, A. I. Virulence and antimicrobial resistance factors of Enterococcus spp. isolated from fecal samples from piggery farms in Eastern Cape, South Africa. BMC Microbiol 15, 136 (2015).
https://doi.org/10.1186/s12866-015-0468-7 |
| [16] | Yuan, T and Pia, Y (2023). Hospital wastewater as hotspots for pathogenic microorganisms spread into aquatic environment: A Review. Frontier in Environmental Science, 10: 17-34. |
[9, 16]
.
Hospitals accumulate large amounts of antibiotics and disinfectant resistant human-related pathogens. For example,
Acinetobacter baumannii,
Citrobacter freundii,
Klebsiella, Aeromonas, Pseudomonas,
S. aureus (
E. coli, P. aeruginosa,
Enterobacter spp, coagulase-negative
Staphylococcus,
Salmonella spp,
Shigella spp., which cause serious hospital-associated infections, have shown resistance to multiple antibiotics
| [15] | Uzoije U. N., Moses I. B., Nwakaeze E. A., Uzoeto H. O., Otu J. O., Egbuna N. R., Ngwu J. N., Chukwunwejim C. R., Mohammed D. I., Peter I. U., Oke B and Iroha I. R (2021). Prevalence of Multidrug-resistant Bacteria Isolates in Waste Water from Different Hospital Environment in Umuahia, Nigeria. International Journal of Pharmaceutical Sciences Review and Research, 69(2): 25-32. |
[15]
.
The abundance of antibiotic resistance in the hospital waste water has become a major global public health concern as a result of the widespread use of antibiotics in healthcare systems. If the resistant bacteria are carrying a transmissible gene, they transfer resistant genes to other community bacteria so that infection caused by these bacteria are usually difficult to treat, and it also decreases the antibiotic pool for the treatment of bacterial infections.
Without suitable treatment, the discharge of antibiotic-resistant bacteria (ARB) in hospital wastewater poses ecological and ARB evolution risks to aquatic environments and humans. This study was designed to isolate, characterize and determine the pattern of resistance of bacteria from hospital waste water effluents to various antibiotics.
2. Material and Methods
2.1. Study Area
The study was carried out at Alex Ekwueme Federal Teaching Hospital Abakaliki, Ebonyi State Abakaliki. Abakaliki is the capital city of Ebonyi State, located 64 kilometers southeast of Enugu in southeastern Nigeria. The majority of the residents are Igbo rice and salt producers. The metropolis of Abakaliki is located in the eastern section of the country at latitude 6.3°E and longitude 8.1°N
| [2] | Adibe-Nwafor, J. O., Uduku, N. D., Iroha, C. S., Ibiam, F. A., Onuora, A. L., Nwafor, K. A., Peter, I. U., Iroha, I. R (2023). Distribution and Antibiotic Resistance Profile of Extended Spectrum Beta-Lactamase Producing Escherichia coli from Fish Farms within Abakaliki Metropolis. Advance in Research, 24(5): 175-184. |
[2]
.
2.2. Ethical Clearance
Ethical clearance (NHREC/16/05/22/361) was collected from the research and ethics committee of Alex Ekwueme Federal Teaching Hospital, Abakaliki, Ebonyi State, before the commencement of sample collection. Every fundamental study was done in accordance with the ARRIVE guidelines.
2.3. Sample Collection
A total of Two hundred and fifty (250) wastewater samples was collected from AEFUTHA at different intervals as follows:
1) From the untreated wastewater outlet pipe of the hospital before it enters the sewer system,
2) Sewage treatment plant (using activated sludge), and
3) Treated water (effluent) before being discharged.
A volume of 250ml wastewater was collected from each site using plastic bottles (sterilized with 70% ethanol for 3 min followed by three times rinsing with sterile distilled water). The collected samples were preserved on ice, and transported to the microbiology laboratory unit of Ebonyi State University for microbiological analysis.
2.4. Bacterial Isolation from Treated Hospital Wastewater
The HWW samples were grouped and pooled then thoroughly shaken to get a homogeneous mixture before a portion was taken for culture. Serial 10-fold dilutions of wastewater samples were prepared in 9ml volume of sterile waster. The bacteria were cultivated by streak plating 0.5 ml of each of the desired serial dilutions of suspensions, 7th and 8th (10−7 and 10−8) dilutions of HWW. Duplicate samples were plated onto MacConkey (MAC) agar, Mannitol Salt agar (MSA), Salmonella/Shigella agar and Cetrimide agar and then incubated at 37°C for 24 hours. After obtaining pure colonies and recording important features, isolated bacteria were further identified at the species level by biochemical tests. The obtained pure cultures of the isolates were maintained at 4°C as agar slants and as glycerol stocks at −20°C in the same media broth containing 25% glycerol
| [8] | Iroha, I. R., Orji, J. O., Onwa, N. C., Nwuzo, A. C., Okonkwo, E. C., Ibiam, E. O., Nwachi, A. C., Afuikwa, F. N., Agah, V. M., Ejikeugwu, E. P. C., Agumah, N. B., Moses, I. B., Ugbo, E., Ukpai, E. G., Nwakaeze, E. A., Oke, B., Ogbu, L and Nwunna, E (2019). Microbiology Practical Handbook. (Editor; Ogbu. O), 1st Edition. Charlieteximage Africa (CiAfrica Press), Pp: 344. |
[8]
.
2.5. Morphological Identification and Biochemical Identification
Morphological and biochemical identification of Bacteria were carried out using Gram stain, Motility test (Hanging drop), Catalase, Coagulase and Carbohydrate fermentation tests.
2.6. Preparation of 0.5 McFarland Turbidity Equivalent Standards
Turbidity standard equivalent of 0.5 McFarland standards was prepared by adding 1.0 ml of concentrated tetraoxosulphate (VI) acid (H
2S0
4) to 99 ml of distilled water; then 0.5 g of dehydrated barium chloride (Bacl2. H2O) was dissolved in 50ml of distilled water in a separate flask. A volume of 0.6ml of barium chloride solution was introduced into tetraoxosulphate (vi) acid solution (99.4 ml) in a separate conical flask. It was mixed well to obtain 0.5 MacFarland turbidity equivalent standards. Some portions of the mixed solutions were transferred to test tube and stored at room temperature (28°C). This was used to compare the turbidity of the test organisms prior to susceptibility test
| [7] | Ilang D. C., Peter I. U., and Iroha I. R (2023). Antibiotic Resistance Profile of Clinical Importance Biofilm forming Extended Spectrum Beta-lactamase and Carbapemase Phenotype in Gram-negative bacteria isolates. International Journal of Pharmacognosy and Life Science, 4(2): 120-127. |
[7]
.
2.7. Standardization of Test Bacteria
All the test bacteria were standardized before use by inoculating a loopful of a 24 hrs culture of the test organism from a nutrient agar slant into test tubes with 5ml sterile water. It was diluted using loopful of the test organism and sterile water in order to obtain microbial population of 1 x 106 colony forming unit per milliliter (cfu/ml) by comparing it with 0.5 MacFarland turbidity standards.
2.8. Antibiotic Susceptibility Testing
This was aseptically carried out using Kirby-Bauer disk diffusion method, and in conformity to the recommended standard of Clinical and Laboratory Standard Institute (CLSI, 2019). A suspension of the test bacteria was made from a 24 hours growth of the test organisms in sterile water to match the 0.5 McFarland turbidity standard. This was seeded on the entire surface of solidified Mueller- Hinton agar (Sigma-Aldrich, U-S-A) plate. The following antibiotic discs with their potencies were used: Oxacillin (OX-30ug), Ofloxacin (OFX-5ug), Lincomycin (L-15ug), Clindamycin (DA-2ug), Vancomycin (VA-10ug), Imipenem (IPM-10ug), Erythromycin (E-5ug), Meropenem (MEM-10ug), Cefoxitin (FOX-30ug), Azithromycin (ATM-30ug), Ciprofloxacin (CIP-5ug), Tetracycline (TE-30ug), Ceftriazone (CRO-30ug), Trimethoprim/Sulphamethoxazole (SXT-25ug), Nalidixic acid (NA-30ug), Nitrofurantoin (F-100ug), Amoxicillin/Clavulanic acid (AMC-30ug), Ceftazidime (CAZ-30ug), Gentamycin (CN-30ug), Ceftriazone (CRO-30ug), Ceftazidime (CAZ-30ug).
The Mueller-Hinton agar (Sigma-Aldrich, U-S-A) plates was incubated at (35°C -37°C) in an aerobic atmosphere for 18-24 hours, after which the percentage susceptibility and resistance were interpreted from the inhibition zone diameters (IZD) produced by the antibiotic discs against the test isolates
| [3] | Clinical and Laboratory Standards Institute (CLSI). Performance standards for antimicrobial susceptibility testing; twenty-eighth edition (M100). Wayne, PA: Clinical and Laboratory Standards Institute; 2019. |
[3]
.
2.9. Determination of Multi-Drug Resistant Index (MDRI)
Multi-drug resistant index (MDRI) was determined to ascertain the resistance level of the isolates which is the number of antibiotics the test isolates are resistant to. The MDRI formular used is:
Where, a = number of antibiotics to which the isolate is resistant to.
b = total number of antibiotics to which the isolates are subjected to.
Test for MRSA (Brilliance MRSA Chromagar): A colony of an organism suspected to be MRSA was smeared and streaked out on this media, along with a known MRSA colony and incubated at 35°C for 24 hours. Thereafter the culture plate was observed and MRSA colonies displayed deep blue coloration (Brilliance blue).
2.10. ESBL Confirmation Using Double Disc Synergy Test (DDST)
Using a double disc synergy test, the
E. coli isolates resistant to the 2
nd and 3
rd generation cephalosporin were further investigated for extended-spectrum beta-lactamase. This was performed on a Mueller Hinton agar with 3 major antibiotics to detect extended-spectrum beta-lactamase-producing organisms. The antibiotics used were ceftazidime (CAZ) 30 µg, cefotaxime (CTX) 30 µg, and amoxicillin-clavulanic acid (AMC) 30 µg, which were placed in a parallel form at a distance of 15 mm from each other with amoxicillin clavulanic acid being at the center of the Petri dish. The culture plates containing the suspected ESBL-producing
E. coli were impregnated with the ceftazidime (CAZ-30ug), Cefotaxime (CTX-30ug) and Amoxicillin-Clavulanic acid (AMC-30ug) and incubated at 37°C for 18-24 hrs. The isolates were considered to be ESBL producers when there was a synergy between the center disc and the two discs with an increase in zone diameter of ≥5 mm for any of the antibiotics tested in combination with the clavulanic acid than when tested alone
| [5] | Ekuma, P. U., Ibiam, F. A., Ekuma, M I., Iroha, C. S., Peter, I. U and Iroha, I. R (2023). Evaluating the Bacteria Profile and Drug Susceptibility Patterns of Urinary Tract Infectious Pathogens in Pregnant Women in Abakaliki Metropolis, Nigeria. International Journal of Pathogen Research, 12(5): 52-62. |
[5]
.
3. Results
Results from our study show that four bacteria were isolated. These were confirmed by various morphological and biochemical tests as shown in
Table 1. Bacteria isolated include
S. aureus, and
E,coli. Some samples yielded no growth.
Escherichia coli presented with the highest frequency (43.24%) of Bacteria isolated followed by
S. aureus (40.54%),
Klebsiella (8.11%) and
Campylobacter (8.11%) as shown in
Table 2. Waste water effluents from the ward complex had the highest distribution of bacteria isolated from this study (40.54%) followed by effluents from the laboratory complex. Effluents from the theatre had the least distribution (2.70%) as shown in
Table 3.
Imipenem was the most resisted antibiotics by
S. aureus (100%) and Ofloxacin was the most effective with distribution of 73.34% each as shown in
Table 4. Nalidixic acid, ceftriazone and Trimethoprim were all completely resisted by
Campylobacter and Gentamicin was the most effective
Table 5. Ceftriazone was the most resisted antibiotics by
Klebsiella (100%). Aztreomicin and Gentamicin were highly effective as shown in
Table 6. Nalidixic acid was the most resisted antibiotics by
E. coli (81.25%). Ceftriazone was the most effective (68.75%) in
Table 7. Multi-antibiotic resistance index of
S. aureus showed the highest values to be from the ward complex and laboratory complex. The least MARI was recorded with
S. aureus isolates from Accident and Emergency as shown in
Table 8. Multi-antibiotic resistance index of
Campylobacter species showed the highest values to be from the ward complex and the least from laboratory complex as shown in
Table 9. Multi-antibiotic resistance index of
Klebsiella species showed the highest values to be from accident/emergency followed by the laboratory complex. The least MARI was recorded with
Klebsiella isolated from the ward complex as shown in
Table 10. Multi-antibiotic resistance index of
E. coli showed the highest values to be from the ward complex and laboratory complex. The least MARI was recorded with
E. coli isolates from the Theatre as shown in
Table 11. Of 15 isolates obtained from this study, 46.66% were MRSA positive and 53.33% were MRSA negative as shown in
Table 12. Out of 16
E. coli isolated from this study, 31.25% were positive for ESBL production while 68.75% were negative for ESBL production as shown in
Table 13.
Table 1. Showing Morphological and Biochemical characteristics of bacteria isolated.
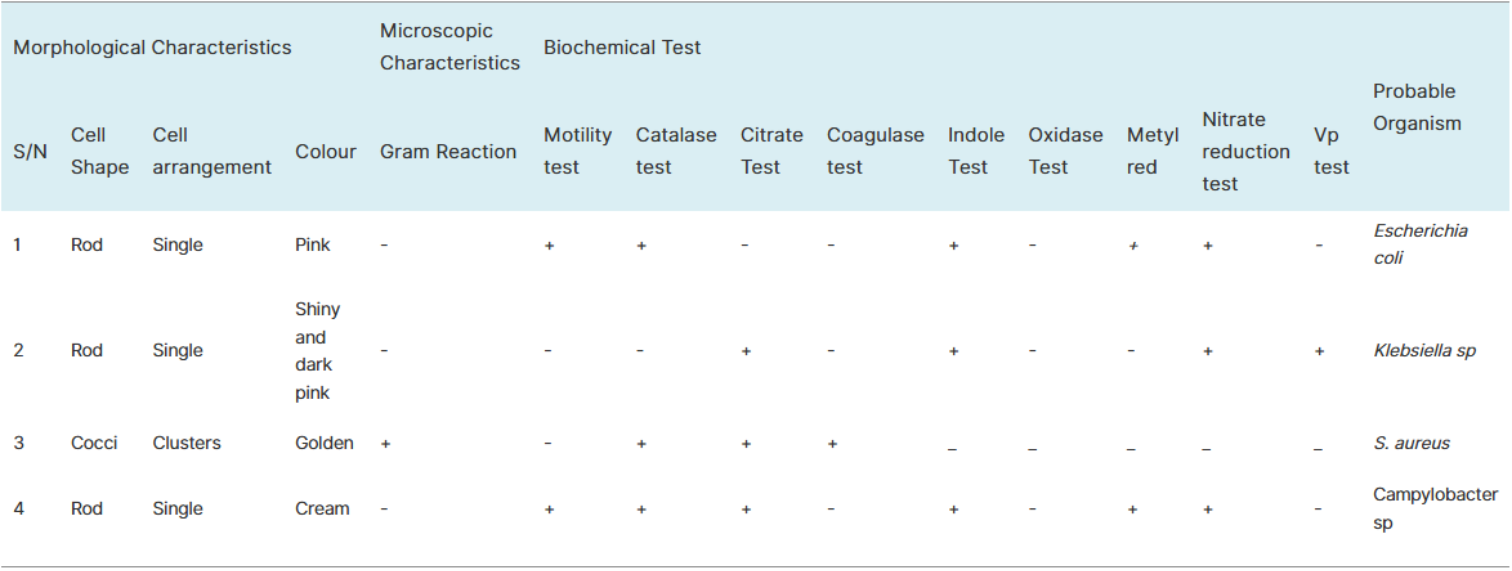
Table 2. Percentage Distribution of Bacteria Isolated from Waste Water in Ae-Futha.
Bacteria Isolated | No Isolated (%) |
Klebsiella sp | 3 (8.11) |
Staphylococcus aureus | 15 (40.54) |
Campylobacter species | 3 (8.11) |
Escherichia coli | 16 (43.24) |
Total | 37 (100) |
Table 3. Percentage Distribution of Bacteria from Various Locations of Ae-Futha.
Locations | No of Bacteria Isolated (%) |
Wards | 15 (40.54) |
Laboratory | 13 (35.14) |
Theatre | 1 (2.70) |
Accident and Emergency | 3 (8.11) |
GOPD | 5 (13.51) |
Total | 37 (100) |
Table 4. Percentage Frequency of Antibiogram of S. Aureus Isolates from Waste Water.
Antibiotics | No of Isolates tested | No Resistant (%) | No Susceptible (%) |
OFX | 15 | 4 (26.66) | 11 (73.34) |
L | 15 | 12 (80.00) | 3 (20.00) |
DA | 15 | 11 (73.34) | 4 (26.66) |
VA | 15 | 8 (53.33) | 7 (46.67) |
IPM | 15 | 15 (100.00) | 0 (0.00) |
E | 15 | 5 (33.33) | 10 (66.67) |
MEM | 15 | 7 (53.33) | 8 (46.67) |
Key: OFX: Ofloxacin, L; Lincomycin, DA: Clindamycin, VA: Vancomycin, IPM: Imipenem, E: Erythromycin, MEM: Meropenem
Table 5. Percentage Frequency of Antibiogram of Campylobacter Sp. Isolates from Waste Water.
Antibiotics | No of Isolates tested | No Resistant (%) | No Susceptible (%) |
ATM | 3 | 1 (33.33) | 2 (66.67) |
F | 3 | 0 (0.00) | 3 (100.00) |
CN | 3 | 0 (0.00) | 3 (100.00) |
TE | 3 | 2 (33.33) | 1 (66.67) |
GR | 3 | 3 (100.00) | 0 (0.00) |
CRO | 3 | 3 (100.00) | 0 (0.00) |
SXT | 3 | 3 (100.00) | 0 (0.00) |
NA | 3 | 3 (100.00) | 0 (0.00) |
Key: ATM; Azithromycin, F: Nitrofurantoin, CN; Gentmicin, TE: Tetracycline, CRO: Ceftriazone, SXT: Trimethoprim, NA: Nalidixic acid
Table 6. Percentage Frequency of Antibiogram of Klebsiella Sp. Isolates from Waste Water.
Antibiotics | No of Isolates tested | No Resistant (%) | No Susceptible (%) |
ATM | 3 | 0 (0.00) | 3 (100.00) |
F | 3 | 0 (0.00) | 3 (100.00) |
CN | 3 | 0 (0.00) | 3 (100.00) |
TE | 3 | 1 (33.33) | 2 (66.67) |
GR | 3 | 1 (33.33) | 0 (0.00) |
CRO | 3 | 3 (100.00) | 0 (0.00) |
SXT | 3 | 2 (66.67) | 1 (33.33) |
NA | 3 | 2 (66.67) | 1 (33.33) |
Key: ATM; Azithromycin, F: Nitrofurantoin, CN; Gentmicin, TE: Tetracycline, CRO: Ceftriazone, SXT: Trimethoprim, NA: Nalidixic acid
Table 7. Percentage Frequency of Antibiogram of Escherichia Coli Isolates from Waste Water.
Antibiotics | No of Isolates tested | No Resistant (%) | No Susceptible (%) |
ATM | 16 | 10 (62.50) | 6 (37.50) |
F | 16 | 7 (43.75) | 9 (56.25) |
CN | 16 | 11 (68.75) | 5 (31.25) |
TE | 16 | 8 (50.00) | 8 (50.00) |
GR | 16 | 12 (75.00) | 4 (25.00) |
CRO | 16 | 5 (31.25) | 11 (68.75) |
SXT | 16 | 8 (50.00) | 8 (50.00) |
NA | 16 | 13 (81.25) | 3 (18.75) |
Key: ATM; Azithromycin, F: Nitrofurantoin, CN; Gentmicin, TE: Tetracycline, CRO: Ceftriazone, SXT: Trimethoprim, NA: Nalidixic acid
Table 8. Multi-Antibiotic Resistance Index of Staphylococcus aureus Isolated from Waste Water Effluents from Various Units of Ae-Futha.
Location of isolate | MARI |
Ward Complex | 1.00 |
Laboratory Complex | 1.00 |
Accident/ Emergency | 0.29 |
Outpatient Department | 0.71 |
Table 9. Multi-Antibiotic Resistance Index of Campylobacter Species Isolated from Waste Water Effluents from Various Units of Ae-Futha.
Location of isolate | MARI |
Ward Complex | 0.75 |
Laboratory Complex | 0.50 |
Table 10. Multi-Antibiotic Resistance Index of Klebsiella Species Isolated from Waste Water Effluents from Various Units of Ae-Futha.
Location of isolate | MARI |
Ward Complex | 0.63 |
Laboratory Complex | 0.75 |
Accident/ Emergency | 0.88 |
Table 11. Multi-Antibiotic Resistance Index of Escherichia Coli Isolated from Waste Water Effluents from Various Units of Ae-Futha.
Location of isolate | MARI |
Ward Complex | 1.00 |
Laboratory Complex | 1.00 |
Accident/ Emergency | 0.63 |
Outpatient Department | 0.88 |
Theatre | 0.13 |
Table 12. Distribution Of MRSA.
MRSA | No of Isolates (%) |
MRSA Positive | 7 (46.66) |
MRSA Negative | 8 (53.33) |
TOTAL | 15 (100) |
Table 13. Distribution of ESBL producing E. coli.
ESBL | No of Isolates (%) |
ESBL Positive | 5 (31.25) |
ESBL Negative | 11 (68.75) |
TOTAL | 16 (100) |
4. Discussion
Hospital wastewater, a critical yet often overlooked component in the spread of AMR, serves as a signifi cant reservoir and conduit for antimicrobial-resistant bacteria (ARB). Hospital wastewater is a hotspot for ARB due to its high concentration of antibiotics and other pharmaceuticals, which significantly contribute to the development of antimicrobial resistant pathogenic bac teria, and spreading infections globally.
Findings from our study showed heterotrophic bacterial count that generally ranged from between 1.12 x 10
2 cfu to 5.6 x 10
2 cfu. The highest ranges of heterotrophic counts were recorded with our findings from waste water effluents collected from the ward complex and the Laboratory complex. Effluents from the ward complex recorded a range between 3.2 x 10
2 cfu to 4.2 x 10
2 cfu and effluents from the Laboratory complex recorded a range between 2.0 x 10
2 cfu to 5.6 x 10
2 cfu. Effluents from the accident and emergency complex recorded a range between 1.48 x 10
2 cfu to 4.2 x 10
2 cfu. Effluents from the General outpatient department recorded a range between 1.12 x 10
2 cfu to 2.2 x 10
2 cfu. Only one heterotrophic count (1.72 x 10
2) was recorded for the effluents from the theatre complex. Our results disagree with studies by Usman
et al | [14] | Usman, K. M., Arotupin, D. J and Ekundayo, F. O. (2021). Antibiotic resistant pattern of bacteria in untreated hospital wastewaters from Offa Local Government Area, Kwara State, Nigeria. African Journal of Microbiology Research, 15(11), 572-582. |
[14]
who examined waste water from various hospitals in Offa; Kwara State. In their study the mean bacterial count population of wet season samples ranged between 7±4.00 × 10
5 and 150±43.59 × (10
5 cfu/ml), while that of dry season samples ranged between 10±2.00 × 10
5 and 225±67.27 × 10
5 cfu/ml. These variations might be due to differences in study period, strategies of the wastewater disposal system and differences in approaches to infection prevention and control measures.
Findings from our study also show that various bacterial species were isolated from waste water effluents emanating from various sections of the hospital. The bacteria isolated include Staphylococcus aureus, Campylobacter species, Klebsiella and Escherichia coli.
Escherichia coli had the highest distribution (43.24%), followed by Staphylococcus aureus (40.54%), Klebsiella (8.11%) and Campylobacter (8.11%).
Findings from our study are consistent with previous studies from Pakistan
| [13] | Shahzad, A., Mian, A. H., Ul haq, I., Khan, M. A and Ali, K (2021). The emergence of different bacterial pathogens in hospital wastewater samples and their antibiotic resistance pattern. Mater Circular Econ.; 3: 1–8. |
[13]
. Our findings did not agree with findings by
| [10] | Kummerer, K (2004). Resistance in the environment. Journal of Antimicrobial Chemotherapy, 54(2): 311–320. |
[10]
. Their study assessed resistance profile of Bacteria isolated in waste water in a tertiary hospital in Maiduguri. In addition to
E. coli, Klebsiella and
S. aureus, they also isolated
Enterobacter species,
Pseudomonas, Salmonella, Proteus and
Bacillus. Another study in Akure reported the isolation of
E. coli, S. aureus, Bacillus subtilis Pseudomonas aeruginosa, Enterobacter aerogenes, Proteus mirabilis, Salmonella typhi and
Aeromonas hydrophila | [6] | Emoruwa, T., Omoya, F., Okewale, A and Ajayi, K. (2024). Occurrence of extended spectrum beta lactamase (ESBL) producing Escherichia coli in wastewater from two hospitals in Akure. Microbes and Infectious Diseases. |
[6]
.
With respect to location, waste water effluents from the ward complex had the highest distribution of bacterial isolates (40.54%) followed by effluents from the Lab complex (35.14%), General outpatients department (13.51%), Accident and emergency complex (8.11%) and the Theatre complex (2.70%).
Antibiogram results from this study showed that Imipenem was the most resisted of antibiotics by S. aureus (100.00%) and Ofloxacin was the most effective with distribution of 73.34%. Nalidixic acid, ceftriazone and Trimethoprim were all completely resisted by Campylobacter and Gentamicin was the most effective. Ceftriazone was the most resisted antibiotic by Klebsiella (100%) while Aztreomycin and Gentamicin were highly effective. Nalidixic acid was the most resisted antibiotics by E. coli (81.25%) while Ceftriazone was the most effective (68.75%).
Multi-antibiotic resistance index (MARI) of S. aureus showed the highest values to be from the ward complex and laboratory complex. The least MARI was recorded with S. aureus isolates from accident and emergency. Multi-antibiotic resistance index of Campylobacter species showed the highest values to be from the ward complex and the least from laboratory complex. Multi-antibiotic resistance index of Klebsiella species showed the highest values to be from Accident/Emergency followed by the laboratory complex. The least MARI was recorded with Klebsiella isolated from the ward complex. Multi-antibiotic resistance index of E. coli showed the highest values to be from the ward complex and laboratory complex. The least MARI was recorded with E. coli isolates from the Theatre.
Similarly, all bacterial isolates from hospital wastewater collected were found to be 100% resistant to some of the antibiotics tested. Similar observation was reported by Iweriebor
et al. (2015) from Alice, Eastern Cape province of South Africa and European countries. More so, presence of high percentage of drug resistant isolates from hospital wastewater suggests that, hospital wastewater could have contributed massively to the resistances observed among the isolates from the final effluent. Of 15
Staphylococcus aureus isolates obtained from this study, 46.66% were methicillin resistant and 53.33% were not methicillin resistant. A study in Akure reported that 6 out of 13
Staphylococcus isolates obtained from hospital waste water were methicillin resistant
| [6] | Emoruwa, T., Omoya, F., Okewale, A and Ajayi, K. (2024). Occurrence of extended spectrum beta lactamase (ESBL) producing Escherichia coli in wastewater from two hospitals in Akure. Microbes and Infectious Diseases. |
[6]
.
Out of 16
E. coli isolated from this study, 31.25% were positive for ESBL production while 68.75% were negative for ESBL production. Our study disagrees with findings from a study in Ibadan were 13.23% of bacteria isolated were ESBL producers
| [1] | Adenike, O and Ayansina, D (2022). Occurrence of Extended Spectrum Beta Lactamase (ESBL) Producing Gram-Negative Bacteria in Wastewaters from Selected Hospitals in Ibadan, Oyo State, Nigeria. Tropical Journal of Natural Product Research (TJNPR), 6(5), 826-830. |
[1]
. The presence of antimicrobial resistant bacteria from hospital wastewaters showed the spreading of AMR bacteria into the environmental through wastewater. Additionally, the results the present study highlight the importance of infection control measures within hospitals and the prudent antibiotic use to curb the spread of antibacterial resistant bacteria. This finding advocates for the implementation of disinfection of wastewater before discharging to the disposal systems.
The high prevalence of drug-resistant isolates from hospital wastewater samples analyzed suggests their persistence in the hospital environment, and their ability to spread antibiotic resistance due to selection pressure and horizontal gene transfer. Therefore, patients are advised to adhere strictly to the directives of the physician in administration of drugs so as to reduce the cases of antibiotics resistance. Also, adequate liquid waste treatment system should be developed to disinfect pathogens in hospital wastewater effluent before discharging into municipal water supply, so as to prevent diseases associated with hospital wastewater effluent microbes.
The presence of high MDR bacteria in hospital wastewater may impose public health challenges because they can transmit resistance traits to other enteric pathogenic bacteria in the community. Thus, wastewater effluents must be disinfected to minimize the microbial burden at each respective ward or unit. A further study is needed to assess factors that could be associated with bacterial contamination of the sewage system.
Abbreviations
ESBL | Extended Spectrum Beta-lactamase |
AMR | Antimicrobial Resistance |
MDR | Multidrug Resistance |
MDRI | Multidrug Resistance Index |
ARB | Antibiotic Resistance Index |
HWW | Hospital Waste Water |
Author Contributions
Elom Ugochukwu Okpo: Conceptualization, Funding acquisition
Edeson Lucy Ogayi: Data curation, Methodology
Amaechi-Nnaji Victoria Obumneme: Formal Analysis
Odo Ikechukwu Ituma: Investigation
Agumah Nnabuife Bernard: Writing – original draft, Writing – review & editing
Iroha Ifeanyi Romanus: Validation, Supervision
Conflicts of Interest
There was no conflict of interest in this study.
References
| [1] |
Adenike, O and Ayansina, D (2022). Occurrence of Extended Spectrum Beta Lactamase (ESBL) Producing Gram-Negative Bacteria in Wastewaters from Selected Hospitals in Ibadan, Oyo State, Nigeria. Tropical Journal of Natural Product Research (TJNPR), 6(5), 826-830.
|
| [2] |
Adibe-Nwafor, J. O., Uduku, N. D., Iroha, C. S., Ibiam, F. A., Onuora, A. L., Nwafor, K. A., Peter, I. U., Iroha, I. R (2023). Distribution and Antibiotic Resistance Profile of Extended Spectrum Beta-Lactamase Producing Escherichia coli from Fish Farms within Abakaliki Metropolis. Advance in Research, 24(5): 175-184.
|
| [3] |
Clinical and Laboratory Standards Institute (CLSI). Performance standards for antimicrobial susceptibility testing; twenty-eighth edition (M100). Wayne, PA: Clinical and Laboratory Standards Institute; 2019.
|
| [4] |
Dadi, B. R., Girma, E., Tesfaye, M and Seid, M (2021). Assessment of the Bacteriological Profile and Antibiotic Susceptibility Patterns of Wastewater in Health Facilities of Ethiopia. International Journal of Microbiology, 9(6): 45-50.
|
| [5] |
Ekuma, P. U., Ibiam, F. A., Ekuma, M I., Iroha, C. S., Peter, I. U and Iroha, I. R (2023). Evaluating the Bacteria Profile and Drug Susceptibility Patterns of Urinary Tract Infectious Pathogens in Pregnant Women in Abakaliki Metropolis, Nigeria. International Journal of Pathogen Research, 12(5): 52-62.
|
| [6] |
Emoruwa, T., Omoya, F., Okewale, A and Ajayi, K. (2024). Occurrence of extended spectrum beta lactamase (ESBL) producing Escherichia coli in wastewater from two hospitals in Akure. Microbes and Infectious Diseases.
|
| [7] |
Ilang D. C., Peter I. U., and Iroha I. R (2023). Antibiotic Resistance Profile of Clinical Importance Biofilm forming Extended Spectrum Beta-lactamase and Carbapemase Phenotype in Gram-negative bacteria isolates. International Journal of Pharmacognosy and Life Science, 4(2): 120-127.
|
| [8] |
Iroha, I. R., Orji, J. O., Onwa, N. C., Nwuzo, A. C., Okonkwo, E. C., Ibiam, E. O., Nwachi, A. C., Afuikwa, F. N., Agah, V. M., Ejikeugwu, E. P. C., Agumah, N. B., Moses, I. B., Ugbo, E., Ukpai, E. G., Nwakaeze, E. A., Oke, B., Ogbu, L and Nwunna, E (2019). Microbiology Practical Handbook. (Editor; Ogbu. O), 1st Edition. Charlieteximage Africa (CiAfrica Press), Pp: 344.
|
| [9] |
Iweriebor, B. C., Obi, L. C and Okoh, A. I. Virulence and antimicrobial resistance factors of Enterococcus spp. isolated from fecal samples from piggery farms in Eastern Cape, South Africa. BMC Microbiol 15, 136 (2015).
https://doi.org/10.1186/s12866-015-0468-7
|
| [10] |
Kummerer, K (2004). Resistance in the environment. Journal of Antimicrobial Chemotherapy, 54(2): 311–320.
|
| [11] |
Odu, C. E., Egbere, J. O., Onyimba, I. A., Ghamba, P. E., Godiya, S., Isyaka, T. M., Collins- Odu, J. O., Idigo, M. A and Ngene, A. C. (2022). Resistance Profiles of Bacteria Isolated from Wastewater in the State Specialist Hospital Maiduguri. European Journal of Biology and Biotechnology, 3(3), 1–6.
|
| [12] |
Oyeleke, S., Istifanus, N and Manga, S (2008). The effects of hospital solid waste on the receiving environment. International Journal of Integrative Biology, 3(3): 191.
|
| [13] |
Shahzad, A., Mian, A. H., Ul haq, I., Khan, M. A and Ali, K (2021). The emergence of different bacterial pathogens in hospital wastewater samples and their antibiotic resistance pattern. Mater Circular Econ.; 3: 1–8.
|
| [14] |
Usman, K. M., Arotupin, D. J and Ekundayo, F. O. (2021). Antibiotic resistant pattern of bacteria in untreated hospital wastewaters from Offa Local Government Area, Kwara State, Nigeria. African Journal of Microbiology Research, 15(11), 572-582.
|
| [15] |
Uzoije U. N., Moses I. B., Nwakaeze E. A., Uzoeto H. O., Otu J. O., Egbuna N. R., Ngwu J. N., Chukwunwejim C. R., Mohammed D. I., Peter I. U., Oke B and Iroha I. R (2021). Prevalence of Multidrug-resistant Bacteria Isolates in Waste Water from Different Hospital Environment in Umuahia, Nigeria. International Journal of Pharmaceutical Sciences Review and Research, 69(2): 25-32.
|
| [16] |
Yuan, T and Pia, Y (2023). Hospital wastewater as hotspots for pathogenic microorganisms spread into aquatic environment: A Review. Frontier in Environmental Science, 10: 17-34.
|
Cite This Article
-
APA Style
Okpo, E. U., Ogayi, E. L., Obumneme, A. V., Ituma, O. I., Bernard, A. N., et al. (2026). Assessment of Antibiotic-resistant Bacteria in Hospital Waste Water Effluents from a Tertiary Hospital in Abakaliki, Nigeria. Biomedical Sciences, 12(2), 26-33. https://doi.org/10.11648/j.bs.20261202.11
 Copy
|
Copy
|
 Download
Download
ACS Style
Okpo, E. U.; Ogayi, E. L.; Obumneme, A. V.; Ituma, O. I.; Bernard, A. N., et al. Assessment of Antibiotic-resistant Bacteria in Hospital Waste Water Effluents from a Tertiary Hospital in Abakaliki, Nigeria. Biomed. Sci. 2026, 12(2), 26-33. doi: 10.11648/j.bs.20261202.11
 Copy
|
Copy
|
 Download
Download
AMA Style
Okpo EU, Ogayi EL, Obumneme AV, Ituma OI, Bernard AN, et al. Assessment of Antibiotic-resistant Bacteria in Hospital Waste Water Effluents from a Tertiary Hospital in Abakaliki, Nigeria. Biomed Sci. 2026;12(2):26-33. doi: 10.11648/j.bs.20261202.11
 Copy
|
Copy
|
 Download
Download
-
@article{10.11648/j.bs.20261202.11,
author = {Elom Ugochukwu Okpo and Edeson Lucy Ogayi and Amaechi-Nnaji Victoria Obumneme and Odo Ikechukwu Ituma and Agumah Nnabuife Bernard and Iroha Ifeanyi Romanus},
title = {Assessment of Antibiotic-resistant Bacteria in Hospital Waste Water Effluents from a Tertiary Hospital in Abakaliki, Nigeria},
journal = {Biomedical Sciences},
volume = {12},
number = {2},
pages = {26-33},
doi = {10.11648/j.bs.20261202.11},
url = {https://doi.org/10.11648/j.bs.20261202.11},
eprint = {https://article.sciencepublishinggroup.com/pdf/10.11648.j.bs.20261202.11},
abstract = {The study was carried out with an aim to isolate, characterize and determine the antibiotics resistance pattern of bacteria isolated from a tertiary hospital in Abakaliki. Waste water effluents were collected from various units of the Alex Ekwueme University Teaching Hospital, Abakaliki and analysed at the Applied Microbiology Laboratory unit of the Ebonyi State University Abakaliki using standard microbiology techniques. Susceptibility of the bacterial isolates to various classes of antibiotics was determined by disc diffusion method. Production of Extended spectrum beta lactamase was determined by the double disc synergy method and the presence methicillin resistant Staphylococcus aureus was determined using Cefoxitin and methicillin disc diffusion method. Our findings showed that Escherichia coli had the highest frequency of bacterial occurence (43.24%) followed by S. aureus (40.54%), Klebsiella (8.11%) and Campylobacter (8.11%). Waste water effluents from the ward complex had the highest distribution of bacteria isolates (40.54%) followed by effluents from the laboratory complex. Effluents from the theatre had the least distribution (2.70%). Out of 15 S. aureus isolates obtained from this study, 46.66% (7) were MRSA positive while out of 16 E. coli isolated from this study, 31.25% (5) were positive for ESBL production. Multi-antibiotic resistance index of S. aureus showed the highest values to be from the ward complex (1.00) and laboratory complex (0.29). Multi-antibiotic resistance index of Campylobacter species showed the highest values to be from the ward complex (0.75) and the least from laboratory complex (0.50) and that of Klebsiella species showed the highest values to be from accident/emergency (0.88) and the least from the ward complex (0.63). Multi-antibiotic resistance index of E. coli showed the highest values to be from the ward complex and laboratory complex (1.00). The presence of antimicrobial resistant bacteria from hospital wastewaters showed the spreading of AMR bacteria into the environmental through wastewater. The presence of high MDR bacteria in hospital wastewater may impose public health challenges because they can transmit resistance traits to other enteric pathogenic bacteria in the community.},
year = {2026}
}
 Copy
|
Copy
|
 Download
Download
-
TY - JOUR
T1 - Assessment of Antibiotic-resistant Bacteria in Hospital Waste Water Effluents from a Tertiary Hospital in Abakaliki, Nigeria
AU - Elom Ugochukwu Okpo
AU - Edeson Lucy Ogayi
AU - Amaechi-Nnaji Victoria Obumneme
AU - Odo Ikechukwu Ituma
AU - Agumah Nnabuife Bernard
AU - Iroha Ifeanyi Romanus
Y1 - 2026/05/16
PY - 2026
N1 - https://doi.org/10.11648/j.bs.20261202.11
DO - 10.11648/j.bs.20261202.11
T2 - Biomedical Sciences
JF - Biomedical Sciences
JO - Biomedical Sciences
SP - 26
EP - 33
PB - Science Publishing Group
SN - 2575-3932
UR - https://doi.org/10.11648/j.bs.20261202.11
AB - The study was carried out with an aim to isolate, characterize and determine the antibiotics resistance pattern of bacteria isolated from a tertiary hospital in Abakaliki. Waste water effluents were collected from various units of the Alex Ekwueme University Teaching Hospital, Abakaliki and analysed at the Applied Microbiology Laboratory unit of the Ebonyi State University Abakaliki using standard microbiology techniques. Susceptibility of the bacterial isolates to various classes of antibiotics was determined by disc diffusion method. Production of Extended spectrum beta lactamase was determined by the double disc synergy method and the presence methicillin resistant Staphylococcus aureus was determined using Cefoxitin and methicillin disc diffusion method. Our findings showed that Escherichia coli had the highest frequency of bacterial occurence (43.24%) followed by S. aureus (40.54%), Klebsiella (8.11%) and Campylobacter (8.11%). Waste water effluents from the ward complex had the highest distribution of bacteria isolates (40.54%) followed by effluents from the laboratory complex. Effluents from the theatre had the least distribution (2.70%). Out of 15 S. aureus isolates obtained from this study, 46.66% (7) were MRSA positive while out of 16 E. coli isolated from this study, 31.25% (5) were positive for ESBL production. Multi-antibiotic resistance index of S. aureus showed the highest values to be from the ward complex (1.00) and laboratory complex (0.29). Multi-antibiotic resistance index of Campylobacter species showed the highest values to be from the ward complex (0.75) and the least from laboratory complex (0.50) and that of Klebsiella species showed the highest values to be from accident/emergency (0.88) and the least from the ward complex (0.63). Multi-antibiotic resistance index of E. coli showed the highest values to be from the ward complex and laboratory complex (1.00). The presence of antimicrobial resistant bacteria from hospital wastewaters showed the spreading of AMR bacteria into the environmental through wastewater. The presence of high MDR bacteria in hospital wastewater may impose public health challenges because they can transmit resistance traits to other enteric pathogenic bacteria in the community.
VL - 12
IS - 2
ER -
 Copy
|
Copy
|
 Download
Download
